HIGH CONTENT SCREENING AT ULTRASTRUCTURAL LEVEL OF HUMAN CELLS BY HMULTI-CLEM
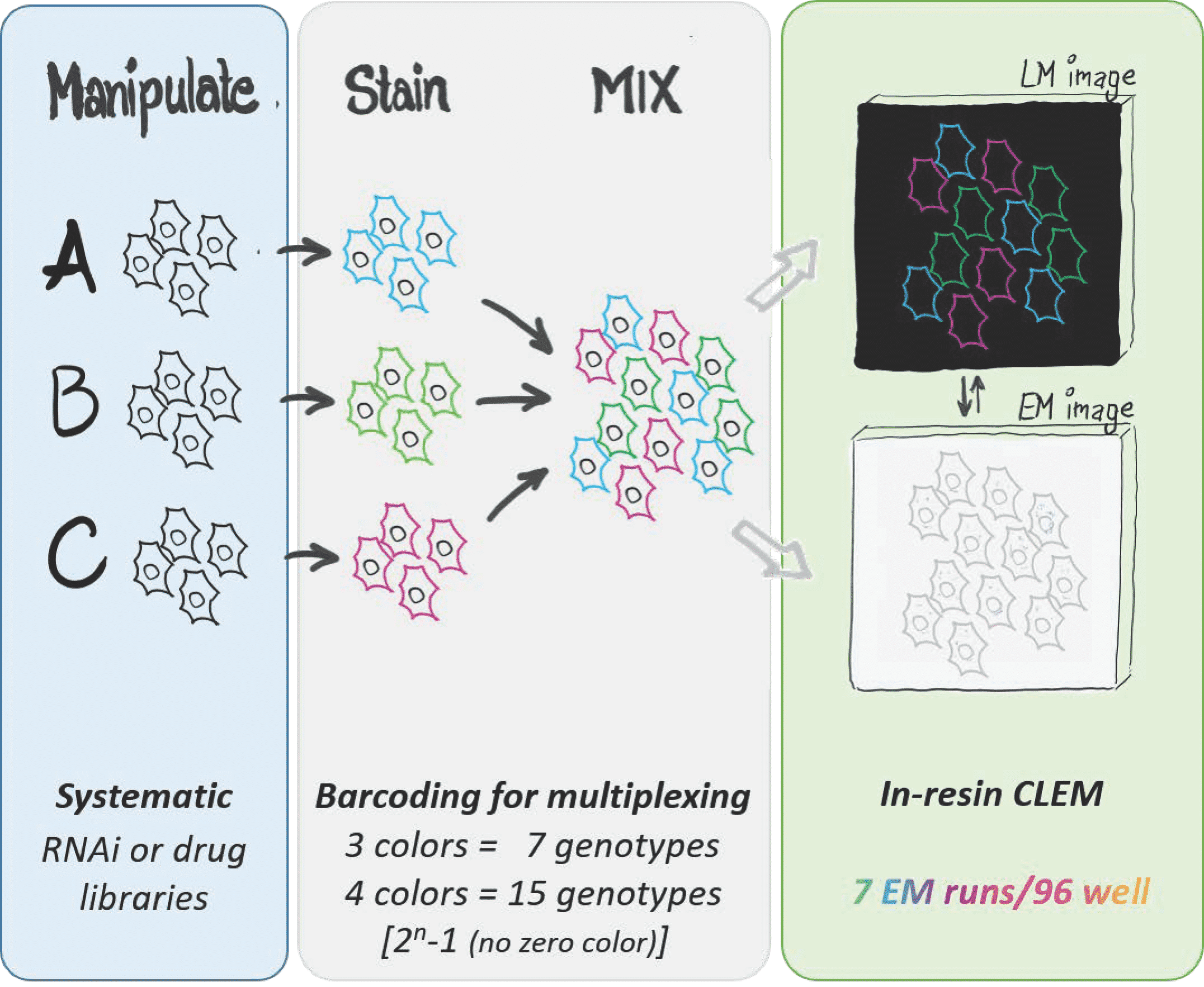
Electron microscopy (EM) imaging of sub-cellular information has, until now, been limited to small experiments due to the lengthy and laborious sample preparation steps. Recently we created a methodology in yeast that enables high-throughput EM analysis - MultiCLEM (Multiplexed Correlative Light and Electron Microscopy). MultiCLEM combines EM with fluorescent barcoding of tens of different cell types for quantitative imaging as mixed population. To distinguish between the barcoded populations, imaging by light microscopy (LM) is correlated to subsequent EM thanks to an established software workflow.
Bringing MultiCLEM to the next level, we have now developed high-throughput ultrastructure screening in human cells – hMultiCLEM. To establish a barcoding protocol universal for a broad variety of cells, we use HeLa (adherent) and Jurkat (non-adherent) as two common human cell lines and compare 4 different approaches to stain their plasma membrane. As a proof of principle, we multiplex 4 different fluorescent dyes that are combinatorially mixed to achieve a total of 15 different barcodes. Using this technology, we have now started the first-ever high content screen at ultrastructural resolution of human cells.
Our approach lays the foundation for expanding systematic screening and high-throughput imaging to the ultrastructural level of human cell lines. With such a capacity, mixed samples of any kind can be analyzed with EM at high speed for diverse applications: time courses, dose responses and quantitative studies. Moreover, sample preparation variations become obsolete as it allows for direct comparison with an internal control. Most importantly, MultiCLEM is built for everyone - it runs on a basic automated widefield microscope, a conventional EM setup, and our freely available software workflow.